Chemokine Receptor-Targeted Library
ChemDiv’s library of small molecule compounds targeting chemokine receptors contains 20,000 compounds.
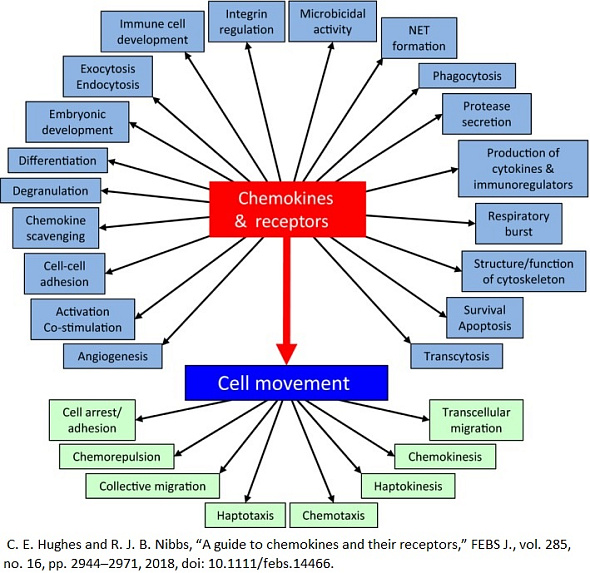
Chemokine receptors are a specific type of cell surface receptors that play a crucial role in the immune system's function by mediating the migration of immune cells toward sites of infection, inflammation, or injury. These receptors bind to chemokines, which are a family of small, secreted proteins. The interaction between chemokines and their receptors triggers a cascade of signals within immune cells, directing their movement (chemotaxis) to areas where their functions are needed. This process is essential for the body's initial defense mechanism against pathogens, the regulation of immune surveillance, and the coordination of immune responses.
The physiological role of chemokine receptors extends beyond the immune system, affecting various normal and pathologic biological processes, including tissue development, angiogenesis, and tumor growth. However, their involvement in disease development is significant, as dysregulated chemokine receptor signaling can contribute to a wide range of diseases. For instance, overexpression or aberrant activation of chemokine receptors can lead to chronic inflammatory diseases, such as rheumatoid arthritis, multiple sclerosis, and asthma, by promoting excessive recruitment of immune cells to tissues, resulting in tissue damage. In cancer, certain chemokine receptors are known to facilitate tumor growth and metastasis by enabling cancer cells to migrate to distant sites and create new tumor growth. HIV uses the chemokine receptor CCR5 as a key entry point into immune cells, highlighting the role of chemokine receptors in infectious disease. Thus, chemokine receptors are critical in health for their role in immune cell trafficking but can contribute to disease pathogenesis when their functions become dysregulated.
The ligand-receptor dynamics within the chemokine superfamily exhibit a remarkable level of complexity. To manage this complexity, the receptors have been categorically divided based on the intricacy of their interactions with ligands, grouping them into distinct classifications. Like all members of the G protein-coupled receptor (GPCR) superfamily, chemokine receptors facilitate signal transduction via specific G-proteins. Despite their morphological similarities to a wide range of other seven-transmembrane segment (7-TMS) receptors, chemokine receptors are distinguished by unique structural features. One notable example is the presence of a specific amino acid sequence, DRYLAIV, within the second intracellular loop domain, underscoring their specialized functional attributes.
Our library of small molecule compounds targeted at chemokine receptors offers significant benefits for drug discovery research, especially given the pivotal role of these receptors in numerous physiological processes and their involvement in a wide array of diseases. This library provides a diverse set of tools to explore the intricate biology of chemokine receptors, facilitating the identification of novel drug targets within this receptor family. By screening these compounds, scientists can uncover specific interactions that modulate immune responses, inflammation, and disease progression, offering insights into new therapeutic strategies.
Targeting chemokine receptors with small molecules allows for the development of highly selective and potent therapeutics with the potential for oral bioavailability and good pharmacokinetic profiles, which are often challenging to achieve with larger biological molecules like antibodies. This selectivity can minimize off-target effects and improve the safety profile of potential drugs.
Furthermore, the versatility of small molecules makes them suitable candidates for the treatment of a broad spectrum of diseases, ranging from inflammatory and autoimmune disorders to cancer and infectious diseases. The ability to modulate chemokine receptor activity with small molecules can lead to the development of drugs that inhibit tumor growth and metastasis, reduce chronic inflammation, or prevent the recruitment of immune cells by pathogens, thereby offering innovative treatments for complex conditions. Overall, our chemokine-targeted small molecule library helps accelerate the drug discovery process, from target identification to the development of candidate drugs, by providing a targeted approach to interfere with disease-related signaling pathways.