Human Kinases Annotated Library
Description
•Kinases are enzymatic proteins that phosphorylate other functional proteins
•Physiologically the kinase enzymatic activity can be modulated by
- Catalytic ATP site binders
- Allosteric binders that cause conformational changes within the catalytic ATP site
•Kinases are well recognized as important therapeutic targets for
- Oncology, e.g. cancer immunotherapy
- Inflammatory diseases, e.g. fibrosis, rheumatoid arthritis
- Cardiovascular system, e.g. cerebral vasospasm, pulmonary arterial hypertension
A unique collection of small molecule compounds with annotated activities for Kinases protein targets
- Annotated activities : 249 kinase targets
- Express Delivery : 640 compounds
- Complete Version : 2585 compounds
Library Composition
Data sources of annotations : Pharos, ChEMBL 25, PubChem, PubMed, Current Patent Literature (CAS, Integrity)
IDNUMBER – ChemDiv Catalog ID (in some instances the same IDNUMBER might have multiple annotation entries due to multiple data sources or because having activity against multiple similar targets);
UNIPROT – SwissProt and ChEMBL Target accesion ID;
Type – character of the measured activity;
Value – Active compounds selection criteria, included only compounds with reported activities < 5 µM;
pubmed_id – PubMed record entry;
doi, patent_id – journal or patent reference to a publication of original data;
For screening data extracted from PubChem, see column assay_description for entry names PUBCHEM_BIOASSAY
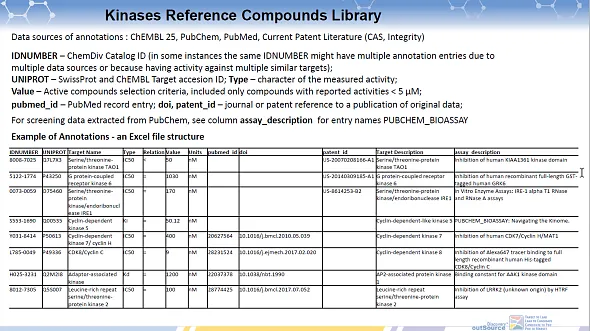
Example of Annotations - an Excel file structure
|
IDNUMBER |
UNIPROT |
Target Name |
Type |
Relation |
Value |
Units |
pubmed_id |
doi |
patent_id |
Target Description |
assay_description |
|
8008-7025 |
Q7L7X3 |
Serine/threonine-protein kinase TAO1 |
IC50 |
< |
50 |
nM |
|
|
US-20070208166-A1 |
Serine/threonine-protein kinase TAO1 |
Inhibition of human KIAA1361 kinase domain |
|
5122-1774 |
P43250 |
G protein-coupled receptor kinase 6 |
IC50 |
= |
1030 |
nM |
|
|
US-20140309185-A1 |
G protein-coupled receptor kinase 6 |
Inhibition of human recombinant full-length GST-tagged human GRK6 |
|
0073-0059 |
O75460 |
Serine/threonine-protein kinase/endoribonuclease IRE1 |
IC50 |
= |
170 |
nM |
|
|
US-8614253-B2 |
Serine/threonine-protein kinase/endoribonuclease IRE1 |
In Vitro Enzyme Assays: IRE-1 alpha T1 RNase and RNase A assays |
|
S553-1690 |
Q00535 |
Cyclin-dependent kinase 5 |
Ki |
= |
50.12 |
nM |
|
|
|
Cyclin-dependent-like kinase 5 |
PUBCHEM_BIOASSAY: Navigating the Kinome. |
|
Y031-8414 |
P50613 |
Cyclin-dependent kinase 7/ cyclin H |
IC50 |
= |
400 |
nM |
20627564 |
10.1016/j.bmcl.2010.05.039 |
|
Cyclin-dependent kinase 7 |
Inhibition of human CDK7/Cyclin H/MAT1 |
|
L785-0049 |
P49336 |
CDK8/Cyclin C |
IC50 |
= |
9 |
nM |
28231524 |
10.1016/j.ejmech.2017.02.020 |
|
Cyclin-dependent kinase 8 |
Inhibition of Alexa647 tracer binding to full length recombinant human His-tagged CDK8/Cyclin C |
|
H025-3231 |
Q2M2I8 |
Adaptor-associated kinase |
Kd |
= |
1200 |
nM |
22037378 |
10.1038/nbt.1990 |
|
AP2-associated protein kinase 1 |
Binding constant for AAK1 kinase domain |
|
8012-7305 |
Q5S007 |
Leucine-rich repeat serine/threonine-protein kinase 2 |
IC50 |
= |
100 |
nM |
28774425 |
10.1016/j.bmcl.2017.07.052 |
|
Leucine-rich repeat serine/threonine-protein kinase 2 |
Inhibition of LRRK2 (unknown origin) by HTRF assay |
Publications
1.J Med Chem 2017 60(14):6337-6352. Discovery of Potent and Selective Inhibitors of Cdc2-Like Kinase 1 (CLK1) as a New Class of Autophagy Inducers. Sun QZ Lin GF Li LL Jin XT Huang LY Zhang G Yang W Chen K Xiang R Chen C Wei YQ Lu GW Yang SY.
2.Bioorg Med Chem Lett 2017 27(11):2617-2621. Developing DYRK inhibitors derived from the meridianins as a means of increasing levels of NFAT in the nucleus. Shaw SJ Goff DA Lin N Singh R Li W McLaughlin J Baltgalvis KA Payan DG Kinsella TM.
3.J Med Chem 2017 60(16):7099-7107. Optimization of Allosteric With-No-Lysine (WNK) Kinase Inhibitors and Efficacy in Rodent Hypertension Models. Yamada K Levell J Yoon T Kohls D Yowe D Rigel DF Imase H Yuan J Yasoshima K DiPetrillo K Monovich L Xu L Zhu M Kato M Jain M Idamakanti N Taslimi P Kawanami T Argikar UA Kunjathoor V Xie X Yagi YI Iwaki Y Robinson Z Park HM.
4.J. Med. Chem. 2015 58(3):1563-1568. A high-throughput screen reveals new small-molecule activators and inhibitors of pantothenate kinases. Sharma LK Leonardi R Lin W Boyd VA Goktug A Shelat AA Chen T Jackowski S Rock CO.
5.ACS Med. Chem. Lett. 2014 5(9):963-967. Hydroxybenzothiophene Ketones Are Efficient Pre-mRNA Splicing Modulators Due to Dual Inhibition of Dyrk1A and Clk1/4. Schmitt C Miralinaghi P Mariano M Hartmann RW Engel M.
6.J. Med. Chem. 2014 57(6):2755-2772. Protein kinase CK-1 inhibitors as new potential drugs for amyotrophic lateral sclerosis. Salado IG Redondo M Bello ML Perez C Liachko NF Kraemer BC Miguel L Lecourtois M Gil C Martinez A Perez DI.
7.Nat. Biotechnol. 2011 29(11):1046-1051. Comprehensive analysis of kinase inhibitor selectivity. Davis MI Hunt JP Herrgard S Ciceri P Wodicka LM Pallares G Hocker M Treiber DK Zarrinkar PP.
8.Bioorg. Med. Chem. Lett. 2011 21(13):4108-4114. Identification of new inhibitors of protein kinase R guided by statistical modeling. Bryk R Wu K Raimundo BC Boardman PE Chao P Conn GL Anderson E Cole JL Duffy NP Nathan C Griffin JH.
9.ACS Med. Chem. Lett. 2011 2(2):154-159. Synthesis and Structure-Activity Relationships of Benzothienothiazepinone Inhibitors of Protein Kinase D. Bravo-Altamirano K George KM Frantz MC Lavalle CR Tandon M Leimgruber S Sharlow ER Lazo JS Wang QJ Wipf P.